This article summarizes "The Fons Constraint: Information-Theoretic Convergence on Encoding Depth in Self-Replicating Systems" (Zenodo, 2026). Full mathematics and code available at the link.
Every biology textbook opens the same way. DNA has four letters: A, T, C, and G. Cells read them three at a time, producing 64 possible three-letter "words" called codons. These 64 codons map to 21 outcomes — 20 amino acids plus a stop signal. This code is universal. Bacteria, mushrooms, blue whales, and you all use it.
Ask a biologist why the code uses triplets — why not doublets (giving only 16 codons) or quadruplets (giving 256) — and you'll usually hear some version of "historical accident." The first self-replicating molecules happened to stumble onto triplets, the story goes, and once life was committed, switching became impossible. It's called the "frozen accident" hypothesis, and it's been the default answer since Francis Crick proposed it in 1968.
The Fons Constraint paper shows that the frozen accident hypothesis is wrong. The genetic code uses triplets because the mathematics of information transfer in noisy channels demands it. Not suggests it. Demands it.
Two Roads to the Same Number
The paper derives the number 64 from two completely independent mathematical frameworks developed decades apart by scientists who never worked on genetics.
The Shannon Path. In 1948, Claude Shannon published the foundational paper of information theory. Among many results, he showed that for any communication channel with a given noise level, there exists an optimal encoding strategy that maximizes the amount of information transmitted per symbol. Applied to a 4-letter alphabet (A, T, C, G) with realistic biological noise levels, Shannon's channel capacity theorem says the optimal codeword length is 3. Shorter codes (length 2, giving 16 options) don't carry enough information per event. Longer codes (length 4, giving 256 options) waste energy on distinctions the receiver can't reliably make. Three is the sweet spot: 4³ = 64.
The Eigen Path. In 1971, chemist Manfred Eigen published his theory of molecular self-replication. He showed that self-replicating molecules face an "error catastrophe" — if the copying error rate is too high relative to the genome length, information is lost faster than selection can maintain it. This sets a maximum viable genome complexity for a given fidelity. Applied to a 4-letter molecular alphabet, Eigen's error threshold constrains viable encoding depth to — once again — three letters per codeword.
Two fields. Two decades. Two completely different mathematical frameworks. One answer: 64.
The Stability Test
A coincidence that works at one noise level isn't very convincing. So we tested the Fons Constraint across nine orders of magnitude of error rate — from ε = 10⁻⁸ (essentially perfect copying) to ε = 10⁻¹ (catastrophically noisy).
The triplet optimum barely moves. Across this enormous range, the optimal encoding depth stays at 3. The genetic code is not perched on a fragile peak that a small perturbation would destroy. It is sitting in a wide, flat valley — the kind of feature that natural selection would find reliably, not the kind it would stumble onto by luck.
This stability is why the code is universal. It's not that the code froze early and couldn't change. It's that there was never anywhere else to go.
What Started With a Wrong Prediction
Honesty compels us to note: the Fons Constraint paper began with a prediction that was wrong.
The original hypothesis was that AI tokenizer vocabulary sizes would cluster near 64, mirroring the genetic code. They don't. Modern AI models use vocabularies of 30,000 to 250,000 tokens — nowhere near 64. This prediction was cleanly falsified.
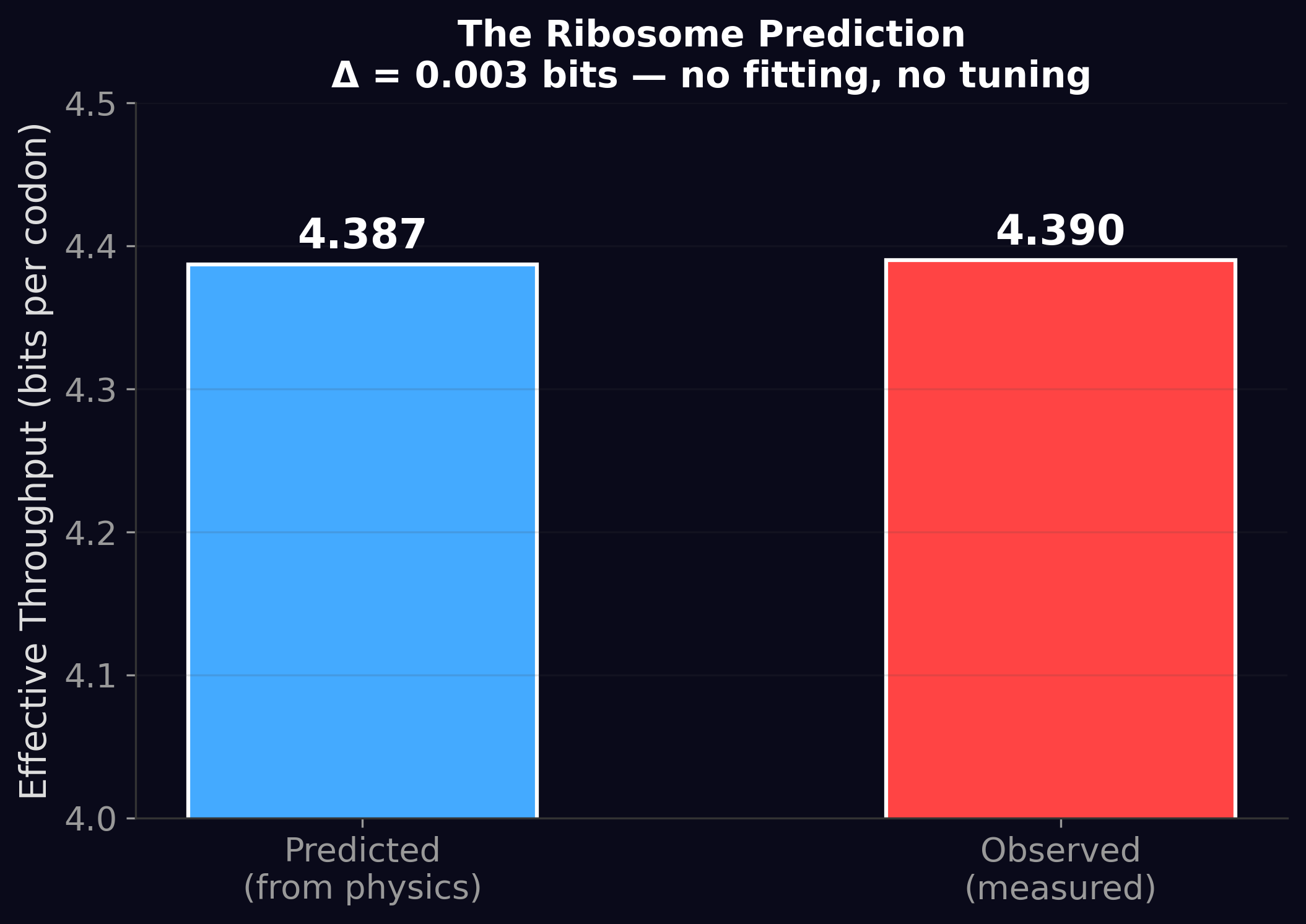
But the falsification revealed something more interesting. While AI vocabularies don't cluster at 64, the effective information per processing event does cluster near the same value as the ribosome (~4.2 bits for AI vs. 4.39 bits for the ribosome). The vocabulary is irrelevant — what matters is the throughput per step. This observation became the seed for everything that followed.
At the Windstorm Institute, we lead with our falsified predictions because that's how science works. The wrong prediction about vocabulary size led directly to the right discovery about throughput constraints. If we had hidden the failure, we would have missed the insight.
What It Means
The Fons Constraint says that the genetic code is not a historical accident frozen in place by contingency. It is a mathematical optimum enforced by the physics of information transfer through noisy channels. Any self-replicating system using a 4-letter alphabet under realistic noise would converge to the same encoding depth.
This has a profound implication for astrobiology: if alien life uses a similar molecular alphabet (four or so distinct chemical letters), the mathematics predicts it would use a similar codon length. The specific chemistry might differ — alien "DNA" might use different molecules entirely — but the information-theoretic architecture would land in the same neighborhood. Physics is the same everywhere. The math doesn't know the difference between adenine and alien-ine.
The Fons Constraint was the first clue that something universal constrains information processing in living systems. The four papers that followed asked: how far does this universality extend? The answer turned out to be: much further than anyone expected.
The arc has since extended beyond biology. Papers 2–6 traced the same throughput constraint through AI tokenization, the neural working-memory limit, and human language. Paper 7 — The Throughput Basin Origin — then asked whether the convergence was architectural, thermodynamic, or data-driven, and found the answer: it’s data-driven. The basin’s width tracks source entropy minus exploitable structure, with no positive evidence of a transformer-specific ceiling. Papers 8 and 9 extended that test to vision, audio, and hardware. The Fons Constraint remains the starting point of the arc.
The Fons Constraint is Paper 1 of the Windstorm series.
Zenodo: doi.org/10.5281/zenodo.19274048 ·
Code & data: github.com/Windstorm-Institute/fons-constraint
Download the full paper (PDF)
Comments & questions
Comments are powered by GitHub Discussions. Sign in with your GitHub account below, or browse all discussions on GitHub. No GitHub account? It takes 30 seconds to make one — or email Grant directly if you'd rather skip it.