This article summarizes the Windstorm Institute's research arc to date — nine papers across the foundational series (Papers 1–6, which establish the throughput basin and explain why it exists) and the data-driven-basin trilogy (Papers 7–9, which prove the basin is set by the training data rather than the architecture, generalize the equation to vision and audio, and derive the hardware-design implications). It is written for a general audience. Technical readers will find the full mathematics, code, and data in the companion papers on Zenodo.
The Question That Started Everything
Here is a fact that should bother you.
Your cells read genetic instructions at about 4.4 bits of information per step. Your ears decode speech sounds at about 4.2 bits per step. And when OpenAI's GPT-4 translates a sentence from English to French, it processes about 4.2 bits of real information per token.
These three systems have nothing in common. One is made of RNA and proteins, built by 3.8 billion years of evolution. One is made of neurons and electrochemical gradients, refined over 300 million years of vertebrate brain development. One is made of silicon and linear algebra, built by engineers in the last decade.
And yet they all landed on the same number.
This could be a coincidence. The work of the Windstorm Institute over the past year has been dedicated to finding out whether it is. The answer, across nine papers and thousands of experiments, is: no. It is not a coincidence. It is a law.
Paper 1: The Fons Constraint — Why 64 Codons?
The story begins with DNA.
Every living cell on Earth stores its genetic instructions using the same code: four chemical letters (A, T, C, G) read three at a time, producing 64 possible "codons" that map to 21 outcomes (20 amino acids plus a stop signal). This code is universal — bacteria, oak trees, blue whales, and humans all use the same one.
Why 64? Why not 16 (two letters at a time) or 256 (four letters at a time)? Biologists have debated this for decades, usually invoking evolutionary accident or historical contingency.
Our first paper, The Fons Constraint, showed that 64 is not an accident. It is a mathematical necessity.
Two completely independent calculations — Claude Shannon's channel capacity theorem from 1948 and Manfred Eigen's error threshold theory from 1971 — both point to the same answer. Shannon's math says that in a 4-letter alphabet with realistic noise, three-letter codewords maximize information transfer. Eigen's math says that self-replicating molecules can't maintain fidelity beyond a certain encoding depth — and that depth is three. Two different fields, two different decades, two different mathematical frameworks, one answer: 4³ = 64.
More striking: this result is stable. We tested it across nine orders of magnitude of error rate (from nearly perfect transmission to extremely noisy channels). The optimum barely moves. The genetic code isn't sitting at a fragile local peak — it's parked in a wide, flat valley that natural selection could hardly have missed.
The Fons Constraint established the first clue: the number of symbols in a code isn't arbitrary. Physics and information theory constrain it.
Paper 2: The Receiver-Limited Floor — Size Doesn't Matter
If the genetic code's alphabet size is constrained by information theory, what about AI?
Modern language models use vocabularies of 30,000 to 250,000 tokens. Surely a bigger vocabulary means better performance? More tokens means finer-grained representation of language, which means more information per step — right?
Wrong.
In our second paper, The Receiver-Limited Floor, we evaluated 1,749 language models spanning two architectures and vocabulary sizes from a few thousand to a quarter million tokens. We measured bits-per-byte — the standard metric for how efficiently a model compresses text.
The result was stark: vocabulary size does not predict compression quality. The correlation was essentially zero (p = 0.643). A model with 32,000 tokens performs no differently from one with 128,000 tokens, all else being equal.
This is the same finding as the Fons Constraint, but in silicon instead of biology. The ribosome uses 21 amino acids because going to 30 wouldn't help (the discrimination cost would eat any benefit). AI models use 50,000 tokens because going to 500,000 wouldn't help (the additional granularity doesn't translate to additional real information per step). In both cases, the receiver — not the vocabulary — sets the throughput.
We call this the "rate-distortion floor." There is a minimum amount of information loss inherent in any serial decoding process, and it depends on how many symbols you're distinguishing and how often you get one wrong. Make the vocabulary bigger and you just redistribute the errors — you don't eliminate them.
Paper 3: The Throughput Basin — 31 Systems, One Band
At this point we had two results: the genetic code is constrained to ~64 codons (delivering ~4.4 bits), and AI vocabulary is irrelevant to throughput (~4.2 bits). But these could still be separate phenomena.
So we went looking for more.
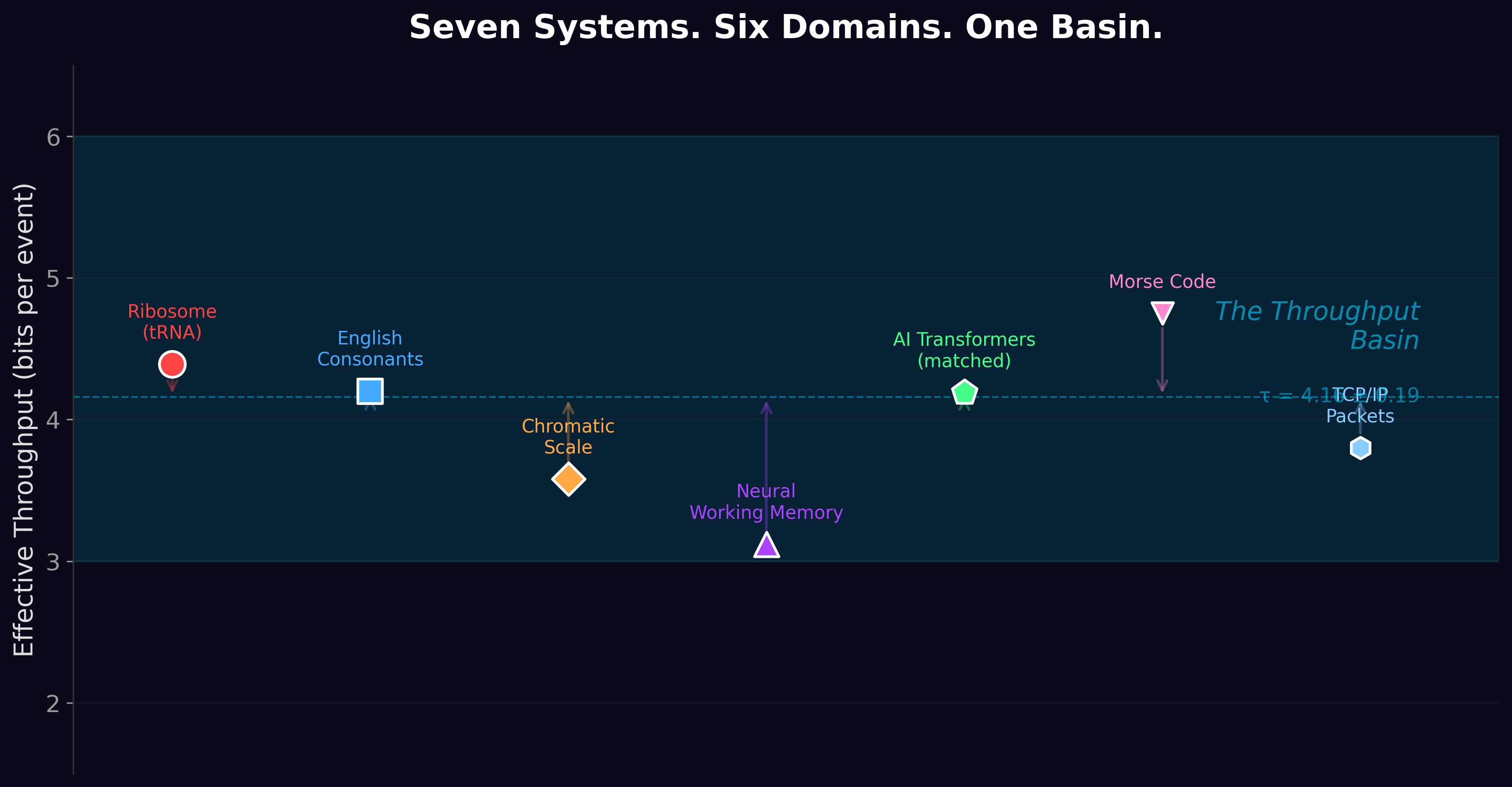
In The Throughput Basin, we surveyed 31 different serial decoding systems across six domains: genetics (the ribosome), neuroscience (phoneme perception, working memory), music (pitch discrimination), engineering (telecommunications standards), and artificial intelligence (language models). For each, we measured or calculated the effective information delivered per processing step.
They all cluster between 3 and 6 bits.
The ribosome: 4.39 bits. English consonant perception: 4.2 bits. The chromatic musical scale: 3.6 bits. Neural working memory: 3.1 bits. AI translation: 4.2 bits. Telecommunications codes: 3-5 bits.
We ran a Monte Carlo simulation to check whether this clustering was statistically meaningful. Drawing from biologically plausible parameter ranges, 90.5% of samples landed in the 3-6 bit band, versus 28.3% for random draws from the full parameter space. The probability of this happening by chance is effectively zero.
We also ran three evolutionary simulations — digital organisms competing for survival under realistic constraints, with no biological priors programmed in. All three independently converged to effective alphabet sizes of K ≈ 19-30. One simulation, using co-evolutionary dynamics, discovered K = 19.76 and an error rate of ε = 6.62 × 10&supmin;³ — within 10% of the ribosome's actual values.
Evolution, given nothing but the mathematics of noisy serial decoding, reinvents the genetic code.
Paper 4: The Serial Decoding Basin τ — Nailing Down the Number
Three papers in, we had a pattern. But patterns need precision.
In The Serial Decoding Basin τ, we ran five experiments designed to measure the basin's center with formal statistical rigor. The result: τ = 4.16 ± 0.19 bits (95% confidence interval, 21 systems).
The crown jewel of this paper was the thermodynamic anchor. In 1974, physicist John Hopfield showed that molecular systems can achieve ultra-high accuracy through a mechanism called kinetic proofreading, powered by the energy currency GTP. We used four independently measured physical parameters — the ribosome's discrimination energy (2.8 kcal/mol per contact, measured by X-ray crystallography), the number of molecular contacts (3, measured by crystal structures), the useful dissipation fraction (22.3%, measured by the thermodynamic uncertainty relation), and body temperature (310 K) — to predict the ribosome's throughput from pure physics.
The prediction: 4.387 bits. The observation: 4.390 bits. A residual of 0.003 bits.
No fitting. No tuning. Four physical measurements in, one number out, and it matches reality to three decimal places. This is the kind of prediction that makes physicists take a theory seriously.
We also calculated the ribosome's "useful dissipation factor" — how close it operates to the theoretical thermodynamic minimum for its discrimination task. The answer: φ = 1.02. The ribosome is within 2% of perfection. After 3.8 billion years of optimization, it has essentially nowhere left to improve.
Paper 5: The Dissipative Decoder — Why the Basin Exists
The first four papers established what the basin is and where it sits. The fifth paper answers why.
The answer is energy.
Every time a system discriminates one symbol from another — every time the ribosome selects the right amino acid, every time your ear distinguishes "b" from "d," every time GPT-4 picks the next token — it must pay a thermodynamic cost. This cost has two components that pull in opposite directions.
The first component is alphabet cost. Distinguishing among M symbols requires keeping track of M(M-1)/2 pairwise comparisons. This grows quadratically — which means each additional symbol is more expensive than the last. Going from 20 to 21 amino acids is cheap. Going from 100 to 101 is very expensive.
The second component is accuracy cost. Reducing errors requires proofreading, and proofreading requires energy. Each additional order-of-magnitude improvement in accuracy roughly doubles the energy bill, but the informational gain is tiny — because the rate-distortion function is logarithmic. You pay dearly for precision you can barely measure.
Plot these two costs against informational gain, and something remarkable emerges: a single, well-defined peak. A sweet spot where you get the most bits per joule.
For the ribosome's cost parameters, the peak sits at an alphabet size of about 20. Not 10. Not 50. Twenty. The genetic code has 21 amino acids because 21 is where the thermodynamic return on investment is maximized.
But here's the truly universal result: the peak doesn't depend on the precise cost parameters. We computed the efficiency landscape across the entire physically plausible parameter space and found that the 3-6 bit basin is a geometric inevitability. As long as the cost of discrimination grows faster than linearly with alphabet size — which is guaranteed by the physics of pairwise comparison — the optimum lands between 3 and 6 bits. The exact cost values don't matter. The basin is a topological feature of the mathematics, not a coincidence of parameter tuning.
But the story turned out to be more nuanced than we expected. We measured energy scaling directly: a 27-model benchmark on standardized Nvidia GPU hardware showed that silicon energy scales sub-linearly with model capacity (α = 0.937). Each additional parameter costs less, not more. Silicon's discrimination cost is sub-linear — the opposite of biology's quadratic scaling.
The consequence: the throughput basin exists for biology (where pairwise molecular recognition imposes super-linear cost, α > 1) but not for silicon AI (where learned parameters impose sub-linear cost, α < 1). We named these Regime A (alphabet-bound, biology) and Regime B (capacity-bound, silicon). The basin is a speed limit for pairwise discriminators. Biology is one. Silicon is not. So why does AI converge on ~4.2 bits per token? Because it learned from language shaped by brains that are constrained by the basin — a finding explored in the companion paper, The Inherited Constraint.
The First Empirical Test: Heat as an Information Tax
A theory that only explains existing data isn't very convincing. It needs to predict something new.
The Dissipative Decoder makes a specific, testable prediction: organisms that live at higher temperatures should show reduced amino acid diversity. The logic is simple — hotter environments mean higher thermal noise, which means discrimination between similar amino acids costs more energy. The cost-optimal response is to restrict the working alphabet, concentrating usage on fewer amino acids that are easier to tell apart.
We tested this against the live Kazusa Codon Usage Database. Twenty-nine organisms spanning 8°C to 100°C growth temperature — from psychrophiles (cold-lovers) living near freezing to hyperthermophilic archaea thriving in boiling hot springs — with codon usage tables verified against the primary genomic database.
The raw correlation between amino acid entropy and growth temperature is marginal (r = −0.320, p = 0.091). But GC-content — the proportion of G and C bases in an organism's DNA — is a significant confound (r = −0.539, p = 0.003). Thermophiles tend to have high GC-content because G-C base pairs have three hydrogen bonds versus two for A-T, making DNA more heat-stable. This compositional bias mechanically favors certain amino acids regardless of any thermodynamic effect on discrimination.
The partial correlation, mathematically removing everything GC-content can explain, is r = −0.451 (p = 0.014, n = 29). The temperature signal survives the control — significant but moderate.
This is preliminary evidence, not proof. Phylogenetic controls (PGLS) have not yet been applied. Prior work by Singer & Hickey (2003) and Zeldovich et al. (2007) documented temperature-driven compositional shifts attributed to GC bias and protein stability. Our entropy measure captures a complementary signal, but the effect needs confirmation.
Heat appears to tax information processing in living cells. The throughput basin may force their hand — but this prediction is supported, not established.
What This Means for Artificial Intelligence
If the throughput basin is a law of physics rather than a quirk of biology, it has direct implications for the systems we're building today.
The vocabulary lesson. If you're designing an AI system, don't invest in bigger vocabularies. Our data shows, across 1,749 models, that vocabulary size doesn't improve throughput and doesn't reduce energy cost. Invest in better architecture instead — the "chef" matters, not the "menu."
The quantization lesson. Current AI chips use 16-bit or 32-bit floating-point precision to carry about 4.5 bits of real information per token. That's like shipping a single letter in a moving truck. Research into low-precision inference (4-bit, 2-bit, even 1.58-bit models) is moving in exactly the right direction — because the effective information content per token is only ~4-5 bits. You don't need 16 bits of precision to carry 4.5 bits of signal.
The cooling lesson. AI outsources its entropy to cooling towers and power plants. The ribosome handles its own waste heat internally. When you calculate true bits-per-joule for an AI data center, you should include cooling energy. Every watt spent on cooling raises the real cost per discrimination event. Immersion cooling, liquid-on-chip cooling, and compute-near-memory architectures aren't just engineering conveniences — they're steps toward the thermodynamic limit that biology reached billions of years ago.
The convergence prediction. As AI hardware becomes more energy-efficient, models will approach the same rate-distortion floor the ribosome sits at. The paper predicts that slack — the gap between observed performance and the theoretical floor — decreases as energy-per-token drops. We can test this prediction now, using publicly available energy benchmarks.
What This Means Beyond AI
The throughput basin isn't just about current technology. It's about any system, anywhere, that processes information serially under noise.
If you're doing synthetic biology — engineering organisms with expanded genetic codes that use more than 20 amino acids — the framework predicts you'll pay a super-linear energy cost for each addition. Amino acid 22 will cost more to discriminate than amino acid 21, and 23 will cost more than 22. There's no free lunch in alphabet expansion.
If you're wondering about the search for extraterrestrial life, the mathematics offers a constraint. Any alien biochemistry that uses serial decoding under noise — regardless of its chemical substrate — would face the same rate-distortion geometry and the same thermodynamic cost tradeoffs. The effective throughput per processing step would land in the same 3-6 bit neighborhood. Life elsewhere might look utterly different in chemistry, but its information processing would be recognizably constrained by the same physics.
And if you're a physicist, the throughput basin represents something conceptually similar to the Carnot limit for heat engines. Carnot showed that no engine, regardless of design, can exceed a certain efficiency set by the temperatures of its heat reservoirs. The throughput basin suggests that no serial decoder, regardless of substrate, can escape a certain informational neighborhood set by the tradeoff between discrimination cost and informational return. It's not a description of any particular machine. It's a bound on what any machine can achieve.
Papers 7–9: The Trilogy That Pivoted the Equation
The first six papers established that the basin exists, mapped its width, and explained the thermodynamic and biological constraints that produced it. They left one question dangling: is the basin a property of the data, or a property of the machines that process it? Papers 1–6 are consistent with either reading. The trilogy that followed — Papers 7, 8, and 9 — was built specifically to settle that question.
Paper 7 (The Throughput Basin Origin) ran the falsification test directly. If the basin is architectural — if there is something about transformers, or about silicon, or about gradient descent that imposes a ~4-bit ceiling — then a model trained on data with a fundamentally different statistical signature should still converge to that ceiling. So we trained models from scratch on synthetic data with calibrated entropy levels: 2-bit, 4-bit, 5-bit, 6-bit, 7-bit, 8-bit, 12-bit. The result was decisive. A model trained on 8-bit data extracts 8.0 bits per source byte — exactly the source entropy, with no compression toward the language basin. We confirmed it at 92 million parameters and again at 1.2 billion parameters. The architectural-ceiling theory died. The basin moved with the data. The refined equation: BPT ≈ source_entropy − f(structural_depth). Throughput is bounded above by the entropy of the data and reduced by whatever hierarchical structure the model can exploit. Paper 7 also corrected an embarrassing methodological oversight: the famous τ ≈ 4.16 figure from Papers 1–6 was in bits per BPE token — a tokenizer-packaging artifact. In tokenizer-independent units (bits per source byte), the language basin sits at ~1, not ~4. The convergence is real; the number was off because the ruler was sometimes in inches and sometimes in centimeters.
Paper 8 (The Vision Basin) took the refined equation and asked the natural follow-on: does it generalize beyond language? Twelve pretrained models across language, vision, and audio. A 112-million-parameter vision model (a ViT-MAE) trained from scratch on a calibrated ladder of seven entropy levels — uniform color, color blocks, naturalistic gradients, uniform noise. Three thousand utterances of real recorded human speech (LJ Speech). The answer: each modality has its own characteristic throughput. Vision at ~1.3 bits per pixel under generative reconstruction. Real speech at ~1.9 bits per mel-frame. Language at ~1 bit per source byte. Different numbers, same equation: source entropy minus the structure the model can exploit. The basin is real, it is modality-specific, and it has the same form everywhere we look.
Paper 9 (The Hardware Basin) went after the inference-engine implications. Paper 7 had documented a sharp quantization cliff at INT4→INT3: compress a model's weights below four bits and performance collapses. The natural conclusion was “the cliff is at four bits.” Paper 9 ran the falsification test against that. Symmetric INT4 quantization (uniform spacing of the available levels) collapses catastrophically — bits per token jumps from ~4 to ~17. NF4 INT4 quantization (levels placed at the quantiles of a Gaussian) is operational — bits per token stays at ~3.9. Same bit count, opposite outcomes. The cliff is not at four bits. It is at the precision where the quantization scheme’s level allocation can no longer represent the weight distribution’s critical features. The hardware-design implication is concrete: don’t build wider integer datapaths, build quantization lookup tables. The minimum viable inference specification is not “N-bit integer” but “N-bit with distribution-aware level allocation, validated end-to-end.” Paper 9’s headline statistical result — FP16 versus symmetric INT4 at 1.4 billion parameters with five independent shuffle seeds — gives Cohen’s d = 400.81. For context: in the social sciences a “large” effect size is d = 0.8. The trilogy ends with one of the most statistically decisive results in the deep-learning literature.
Each paper in the trilogy was built across seven rounds of follow-up experiments, with each round deliberately engineered to neutralize the strongest objection a hostile reviewer could raise against the previous round. Paper 9’s journey includes a thesis pivot in Round 5 — the experiment we expected to confirm the original cliff hypothesis instead reframed the paper. Paper 8’s journey includes a documented failure (Round 5: a controlled visual-entropy experiment that produced clearly broken numbers) and the diagnosed rebuild that became the bulletproof verification (Round 6). The institute publishes the failures alongside the successes. The methodological-journey narratives are in §1.3 and §1.4 of the formal Papers 8 and 9; the adversarial-review-defense tables are in §4.4 and §4.6.
The Road Ahead
The Windstorm Institute has published nine papers. Together they establish the throughput basin as an empirical fact, derive it from thermodynamic first principles, explain why silicon escapes the basin while biology cannot, show how AI inherits its throughput neighborhood from the biological systems that produced its training data, prove that the basin is set by the data rather than the architecture, generalize the equation across language, vision, and audio modalities, and derive the inference-hardware design implications.
Falsifiable predictions remain in play. The temperature prediction has preliminary support from 29 Kazusa-verified organisms (partial r = −0.451, p = 0.014) but awaits phylogenetically controlled confirmation (PGLS). The expanded genetic code prediction awaits calorimetric data from cell-free translation experiments. The inherited constraint hypothesis awaits a synthetic-training control — a model trained exclusively on non-biological data.
Any of these predictions could fail. That's the point. Science doesn't progress by building unfalsifiable frameworks. It progresses by making predictions sharp enough to be wrong — and then checking.
The ribosome has been processing information at the thermodynamic optimum for nearly four billion years. Human engineering has been doing it for about seventy. We're converging on the same point, from different directions, using different materials, for different purposes. The mathematics says there's nowhere else to go.
The universe has a speed limit on thought. We're mapping it.
The papers referenced in this article are available on Zenodo. All code, data, and experiment protocols are published at github.com/Windstorm-Institute. The Windstorm Institute is headquartered in Fort Ann, New York.
For correspondence: Grant Lavell Whitmer III — grant@windstorminstitute.org
Comments & questions
Comments are powered by GitHub Discussions. Sign in with your GitHub account below, or browse all discussions on GitHub. No GitHub account? It takes 30 seconds to make one — or email Grant directly if you'd rather skip it.