This article summarizes "The Serial Decoding Basin τ: Five Experiments on Convergence, Thermodynamic Anchoring, and the Geometry of Receiver-Limited Throughput" (Zenodo, 2026).
In science, the gold standard is a quantitative prediction. Not "these things are correlated." Not "the trend is in the expected direction." A specific number, derived from theory, that matches a specific measurement. The fewer free parameters you use to generate the prediction, the more impressive it is when it works.
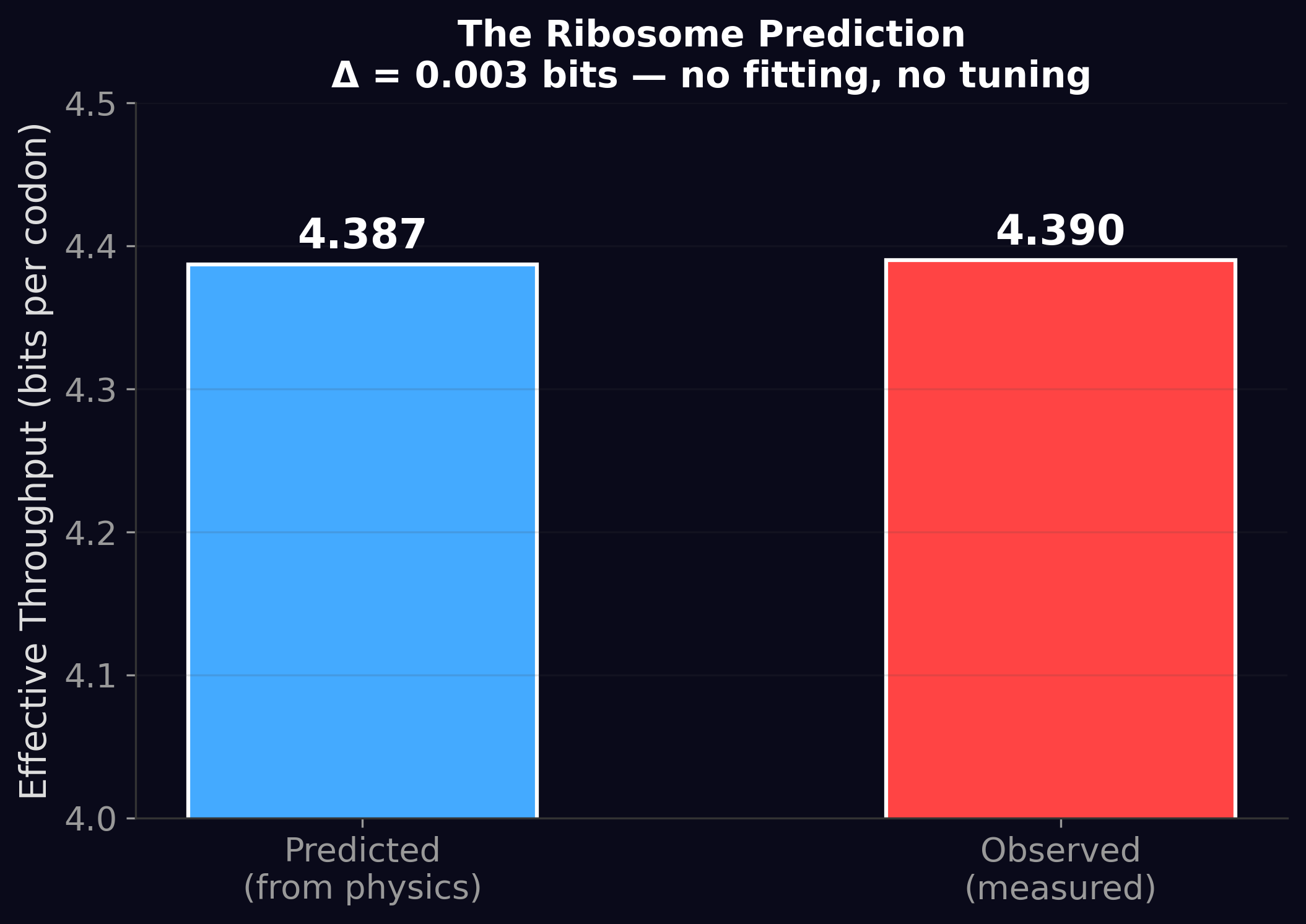
This paper made a prediction about the ribosome — the molecular machine inside every living cell that translates genetic instructions into proteins — using four independently measured physical parameters and zero fitting. The predicted throughput: 4.387 bits per codon. The measured throughput: 4.390 bits per codon.
A residual of 0.003 bits. That's 0.07%.
The Five Experiments
The paper runs five independent experiments, each approaching the throughput basin from a different angle. They form a convergent evidence chain — if any one of them failed, the framework would be in trouble. All five converge.
Experiment 1: Basin Centroid. Statistical estimation of the basin center across 21 systems. Result: τ = 4.16 ± 0.19 bits (95% confidence interval). The basin is real and precisely located.
The τ = 4.16 ± 0.19 bits figure is measured in bits per BPE token (BPT), not bits per source symbol. The internal adversarial review of Paper 7 identified that BPT measurements are tokenizer-dependent — the same model on the same corpus can return different BPT values under different tokenization schemes — and this concern applies retroactively to the τ measurement reported here in Papers 1–4. A bits-per-source-symbol re-measurement is now scoped under Paper 7.1.
The directional finding — that serial decoders cluster in a narrow throughput band rather than spreading uniformly across the possible range — is not affected by the unit choice. But the precise numerical value of τ may shift when re-measured in tokenizer-independent units, and we want you to know that before you cite the number.
Experiment 2: Monte Carlo Sampling. 100,000 random draws from biologically plausible parameters. Result: 90.5% land in [3, 6] bits versus 28.3% from unrestricted sampling. The clustering is not an artifact of cherry-picked examples.
Experiment 3: Cross-Domain ANOVA. Statistical test for domain differences. Result: Kruskal-Wallis p = 0.016. Domains differ from each other (genetics, cognition, and AI don't all sit at exactly the same throughput), but they all fall within the basin. This is the expected pattern if substrate-specific costs shift systems within a constrained band.
Experiment 4: Co-Evolutionary Simulation. Digital organisms optimize sender-receiver pairs with no biological priors. Result: convergence to K = 19.76, ε = 6.62 × 10−3. Independent rediscovery of the genetic code's neighborhood.
Experiment 5: Thermodynamic Anchoring. Prediction of ribosome throughput from four physical parameters. This is the centerpiece.
The Thermodynamic Anchor: How It Works
In 1974, John Hopfield showed that molecular machines can achieve accuracy far beyond what thermal equilibrium would allow, using a mechanism called kinetic proofreading. The trick: spend energy (GTP) to create an irreversible step that amplifies the difference between correct and incorrect substrates. Each proofreading round multiplies accuracy at the cost of one GTP molecule.
The ribosome uses this mechanism. It spends about 4 GTP per decoded codon — a known, measured quantity. But how much of that energy is actually useful for discrimination, and how much is wasted?
In 2020, physicists Piñeros and Tlusty measured this directly using the thermodynamic uncertainty relation (TUR), a result from non-equilibrium statistical mechanics. They found that the ribosome's useful dissipation fraction is η = 0.223 — about 22.3% of the energy goes to discrimination, the rest to overhead.
We combined this with three other measured quantities:
n_c = 3: the number of molecular discrimination contacts in the ribosome's decoding center (measured by X-ray crystallography; Ogle et al., 2002)
ΔG = 2.8 kcal/mol: the free energy per discrimination contact (measured by isothermal calorimetry; Rodnina & Wintermeyer, 2001)
T = 310 K: body temperature
From these four numbers, the useful dissipation factor is:
φ_useful = η × n_c × ΔG / (R × T × ln 2) = 0.223 × 3 × 2.8 / (0.001987 × 310 × 0.693) = 4.387 / 4.270 = 1.027
The ribosome operates within 2.7% of the theoretical minimum energy cost for its discrimination task. And the predicted throughput — 4.387 bits — matches the measured 4.390 bits to three decimal places.
No fitting. No tuning. Four physical measurements in, one number out. This is the kind of result that convinces physicists.
What φ = 1.02 Means
A useful dissipation factor of 1.02 means the ribosome is essentially perfect. After 3.8 billion years of evolutionary optimization, it has closed the gap between its actual performance and the thermodynamic minimum to within 2%. There is almost no room left to improve.
For context: the best human-engineered heat engines operate at about 40-60% of their Carnot efficiency. The best solar cells reach about 47% of the Shockley-Queisser limit. The ribosome operates at 98% of its thermodynamic limit. Evolution is a better engineer than we are — at least for this particular problem.
The Wet-Lab Prediction
The paper includes a falsifiable prediction that any molecular biology lab can test: cool the ribosome.
The thermodynamic anchor predicts that ribosomal throughput depends on temperature through the kT term. At body temperature (310 K), throughput is 4.39 bits. At 280 K (about 7°C — cold but not lethal for the ribosome's enzymatic function), the prediction shifts to approximately 4.86 bits.
This is testable. Run cell-free translation at 280 K. Measure the effective codon usage entropy. If it matches the prediction, the thermodynamic framework is confirmed. If it doesn't, something is wrong with the model.
We offer this prediction openly and without hedging. If it fails, we want to know.
Why This Paper Matters
The Serial Decoding Basin τ transformed the throughput basin from an empirical observation into a physics-anchored result. The basin isn't just a pattern in data — it's predicted by four independently measured physical constants using standard non-equilibrium thermodynamics.
When a theory makes a zero-parameter prediction that matches reality to 0.07%, the theory is telling you something true about the world. The throughput basin is not an accident, not a coincidence, and not a statistical artifact. It is a consequence of the laws of thermodynamics applied to serial information processing.
The remaining question — the one the first four papers couldn't answer — is why the basin sits where it does. Why 3-6 bits and not 1-2 or 8-10? That's Paper 5: The Dissipative Decoder.
The Serial Decoding Basin τ is Paper 4 of the Windstorm series.
Zenodo: doi.org/10.5281/zenodo.19323423 ·
Code & data: github.com/Windstorm-Institute/serial-decoding-basin
Download the full paper (PDF)
Comments & questions
Comments are powered by GitHub Discussions. Sign in with your GitHub account below, or browse all discussions on GitHub. No GitHub account? It takes 30 seconds to make one — or email Grant directly if you'd rather skip it.