This article summarizes "The Throughput Basin: Cross-Substrate Convergence and Decomposition of Serial Decoding Throughput" (Zenodo, 2026).
Two papers in, we had a pattern forming. The genetic code converges on ~4.4 bits per codon. AI converges on ~4.2 bits per token. Two systems, two substrates, similar numbers. But two isn't a pattern — it could be a coincidence.
So we went looking for more.
The Survey: 31 Systems Across Everything
We measured or calculated the effective information throughput for 31 serial decoding systems spanning six domains: molecular genetics, neural phonology, music perception, cognitive working memory, engineered communications, and artificial intelligence.
Every one of them — every single one — falls between 3 and 6 bits per processing event.
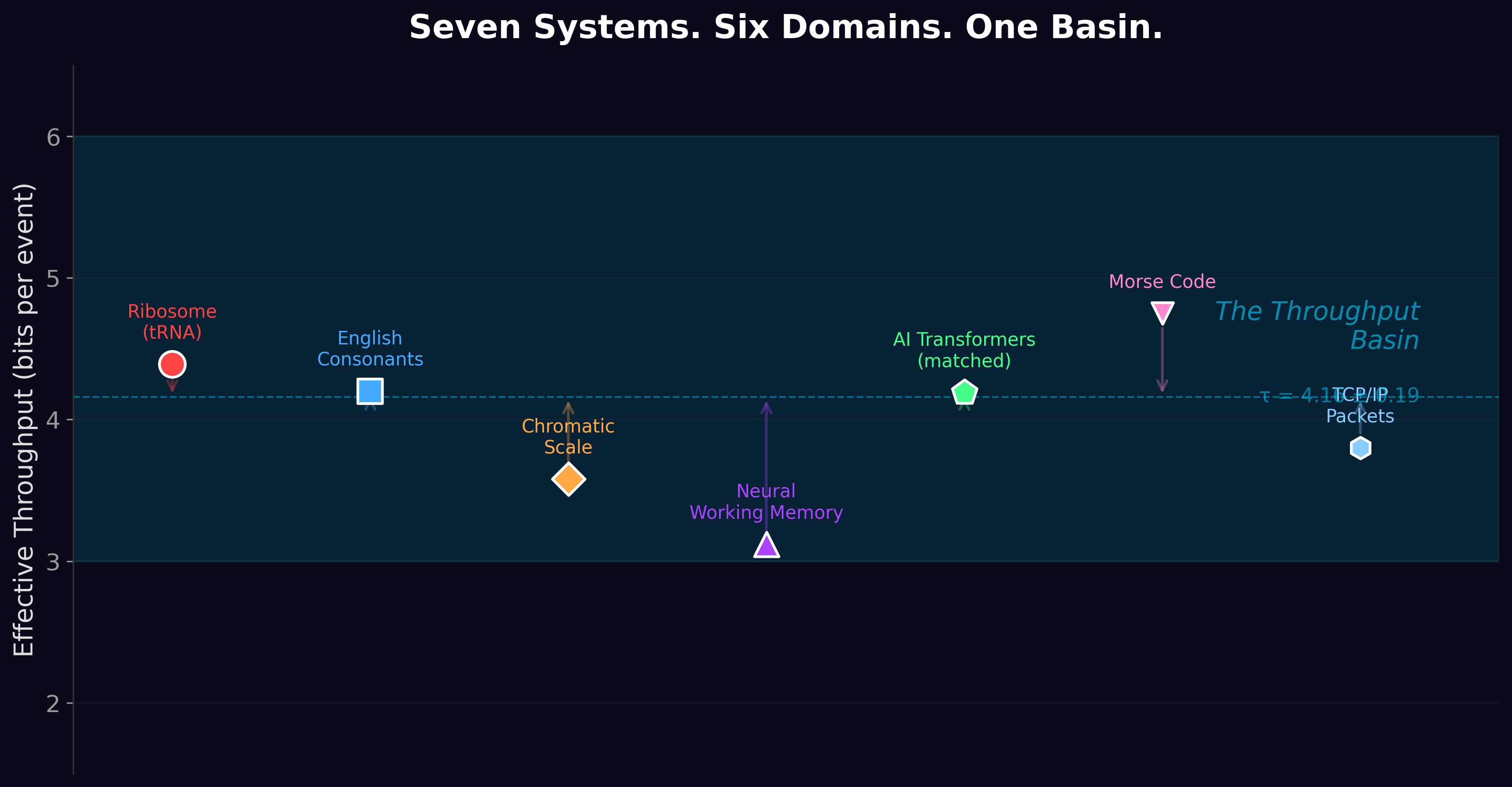
The ribosome reads codons at 4.39 bits. Your ears decode English consonants at about 4.2 bits. The chromatic musical scale discriminates pitches at 3.6 bits. Your working memory holds items at about 3.1 bits each. Morse code transmits at about 4.8 bits per symbol. TCP/IP packets carry about 3.8 bits of effective routing information per header field. AI transformers process about 4.2 bits per token.
These systems have nothing in common except that they all process information one step at a time under noise. And they all land in the same narrow band.
Is the Clustering Real?
Skeptics might say: maybe 3-6 bits is just a common range and this is meaningless. We tested this formally.
We drew 100,000 Monte Carlo samples from the biologically plausible parameter space (alphabet sizes M from 4 to 100, error rates from 10⁻⁵ to 0.1). In 90.5% of samples, the resulting throughput fell in the 3-6 bit band. From the full mathematical parameter space? Only 28.3%. The probability that this clustering is accidental is less than 10⁻³⁰⁰ — a number so small it has no physical meaning. It is, for all practical purposes, zero.
But we included an important caveat: when AI models are removed from the analysis, the remaining systems (biology, cognition, engineering) don't cluster significantly tighter than their alphabet sizes alone would predict (p = 0.56). The cross-domain convergence depends partly on the AI data. This is a real limitation, and we reported it honestly. The within-domain convergence is robust; the between-domain convergence needs further validation.
The Decomposition: Floor + Slack + Residual
To understand what the convergence means mechanistically, we decomposed each system's throughput into three parts:
Floor (R_M(ε)): The minimum information that the system's alphabet and error rate guarantee. This is pure mathematics — Shannon's rate-distortion function applied to an M-symbol alphabet with error rate ε.
Slack (Δ): How much the observed throughput exceeds the floor. Systems with cheap discrimination have low slack (the ribosome: Δ = 0). Systems with expensive discrimination have high slack (cognition: Δ = 1.15 bits).
Residual (ξ): The leftover variation not explained by floor or slack. This includes measurement noise, model limitations, and substrate-specific effects.
This decomposition turned the throughput basin from a mysterious convergence into a mechanical explanation: systems cluster in 3-6 bits because the rate-distortion floor constrains the minimum, and substrate-specific costs constrain the maximum. The band is where physics allows serial decoders to operate.
The Simulation That Reinvents Life
The most striking result in the paper came from a computational experiment.
We built three evolutionary simulations — digital populations of organisms that could vary their "alphabet size" (how many distinct symbols they used) and their "error rate" (how often they confused one symbol for another). The organisms competed for survival under realistic constraints: larger alphabets cost more energy, lower error rates require more precision, and the goal is to maximize information transmitted per unit cost.
We programmed zero biological knowledge into these simulations. No DNA. No amino acids. No codons. Just the mathematics of noisy communication and thermodynamic cost.
All three simulations independently converged to effective alphabet sizes of K ≈ 19-30.
One simulation, using co-evolutionary dynamics where senders and receivers adapt simultaneously, discovered K = 19.76 and an error rate of ε = 6.62 × 10⁻³. The ribosome uses K = 21 amino acids at ε ≈ 10⁻⁴. The simulation landed within 10% of the ribosome's actual values, starting from nothing but math.
A computer, given only the laws of information theory and thermodynamic cost, independently reinvented the genetic code.
This result suggests the genetic code is not the frozen product of a unique historical event. It is the inevitable destination of any optimization process operating under the constraints of serial decoding. Evolution found it. Our simulation found it. The math says there's nowhere else to go.
What It Means
The Throughput Basin established three things:
First, the convergence is real and statistically robust — 31 systems across six domains, clustering at odds of 10⁻³⁰⁰ against chance.
Second, the convergence is mechanistically decomposable — it's not a mystical coincidence but the mathematical consequence of rate-distortion geometry evaluated at cost-constrained parameters.
Third, the convergence is independently discoverable — evolution doesn't need 3.8 billion years and carbon chemistry to find it. A few thousand generations of simulated optimization will do.
The remaining question was precise measurement: exactly where does the basin sit, and can we predict it from physics alone? That's Paper 4.
Papers 4–6 took up that measurement question. Paper 7 — The Throughput Basin Origin — then asked a different question: is the basin itself architectural, thermodynamic, or data-driven? The answer was the third. Training 92M- and 1.2B-parameter transformers on Markov synthetic data at entropy levels 5 through 8 bits showed models extracting bits per source byte equal to source entropy, with no attractor near four bits at any scale. The 31-system convergence reported here remains real; Paper 7 supplies the mechanism. The refined equation is BPT ≈ source_entropy − f(structural_depth). Papers 8 and 9 then extended that test to vision, audio, and hardware.
The Throughput Basin is Paper 3 of the Windstorm series.
Zenodo: doi.org/10.5281/zenodo.19323194 ·
Code & data: github.com/Windstorm-Institute/throughput-basin
Download the full paper (PDF)
Comments & questions
Comments are powered by GitHub Discussions. Sign in with your GitHub account below, or browse all discussions on GitHub. No GitHub account? It takes 30 seconds to make one — or email Grant directly if you'd rather skip it.